VITEK® MS: Healthcare
Microbial Identification in Minutes
VITEK MS is an automated mass spectrometry microbial identification system that uses Matrix Assisted Laser Desorption Ionization Time-of-Flight (MALDI-TOF) technology. VITEK MS contains a comprehensive FDA 510(k) cleared database for bacteria and fungi, including mycobacteria, Nocardia and molds—now including Brucella, Candida auris and Elizabethkingia anophelis.
- Robust & accurate identifications with Advanced Spectra Classifier
- Seamless integration of ID/AST results for optimized workflow
- Complete traceability & flexibility
Fast and Actionable Identification for Better Patient Outcomes
VITEK MS helps laboratories deliver actionable results to clinicians to support informed treatment decisions.
How MALDI-TOF works
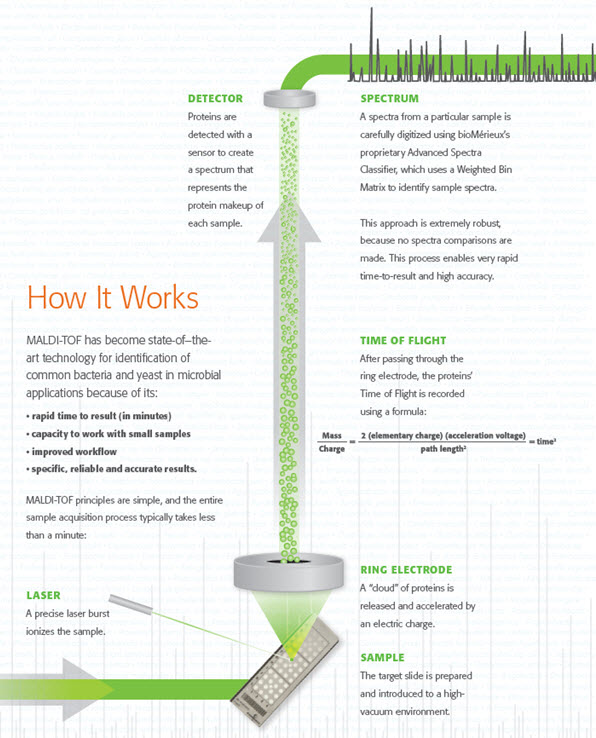
Accurate, Robust Results
bioMérieux’s proprietary Advanced Spectra Classifier offers robust and accurate results needed for optimal patient-care decisions. The VITEK MS system reads each spectrum as a series of peaks that are detected and sorted by mass and intensity. With the use of the Advanced Spectra Classifier, better discrimination is provided as every peak is considered in the calculation of the identification result. There are no spectra comparisons, and the reference spectra are not embedded in the commercial software.
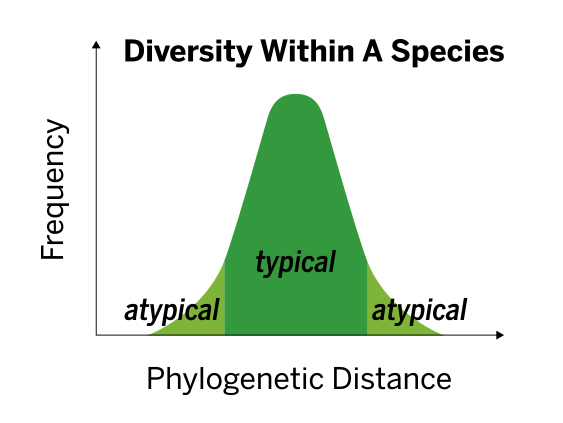
The VITEK MS contains a population built database, accounting for diversity among strains within the same species, leading to robust and accurate performance without the need to modify scores.
The database is built to allow for better discrimination between species. This is accomplished by including spectra for each species generated from:
- An average of 12 strains per species and an average of 35 spectra per species
- Geographical diversity of isolates
- Isolates of different sample origins (e.g., blood, tissue, etc.)
- Isolates subcultured on different media
- Isolates with different incubation times
VITEK MS Knowledge Base V3.2
A comprehensive database update for mycobacteria, Nocardia, and molds—now including Brucella, Candida auris and Elizabethkingia anophelis.
The VITEK MS database is continually expanding to identify new emerging pathogens and clinically relevant species.
The VITEK MS V3.2 FDA 510(k) cleared database for bacteria and fungi contains mycobacteria, Nocardia and molds. This enhanced database includes Mycobacteria tuberculosis (TB), non-tuberculous mycobacteria (NTM), dermatophytes and dimorphic fungi, including the most prevalent species relevant to patient infections. The ability to quickly identify these difficult organisms on a mass spectrometry system allows microbiologists to provide clinicians with actionable results to aid in quicker treatment for diseases such as tuberculosis, osteomyelitis caused by NTM, and serious fungal infections.
VITEK MS’s excellent performance in identifying mycobacteria, Nocardia and molds, allows results to be rapidly released to physicians for better patient care. This helps hospitals save both money (molecular probes; sequencing; send-outs) and time (in-house testing; faster identification of these difficult organisms).
The VITEK MS V3.2 database now includes Brucella, Candida auris, and Elizabethkingia anophelis.
Brucella species are difficult to recognize and hazardous to laboratory personnel. Quick identification reduces exposure to laboratory personnel and enhances safety. Both Candida auris and Elizabethkingia anophelis are new emerging pathogens of clinical significance.
Optimized Workflow and Seamless ID/AST Integration
VITEK MS is designed specifically for optimized workflow in the microbiology laboratory. Its 4-slide capacity enables parallel preparation of samples by four different technologists at separate workstations, with placement on the instrument at the same time. With 48 sample spots per target slide, 192 isolates can be tested per run.
- Guided slide preparation for ID and connection with AST using the VITEK MS Prep Station
- Very few steps to obtain a result – same day ID and AST (with VITEK 2) results
- Less hands-on time to focus on more value added tasks
- Variable throughput for large and small labs (1-192 samples/run)
MYLA®, bioMérieux’s advanced middleware solution, offers real-time reporting and seamless integration of the ID and AST results from VITEK MS and VITEK 2.
 |
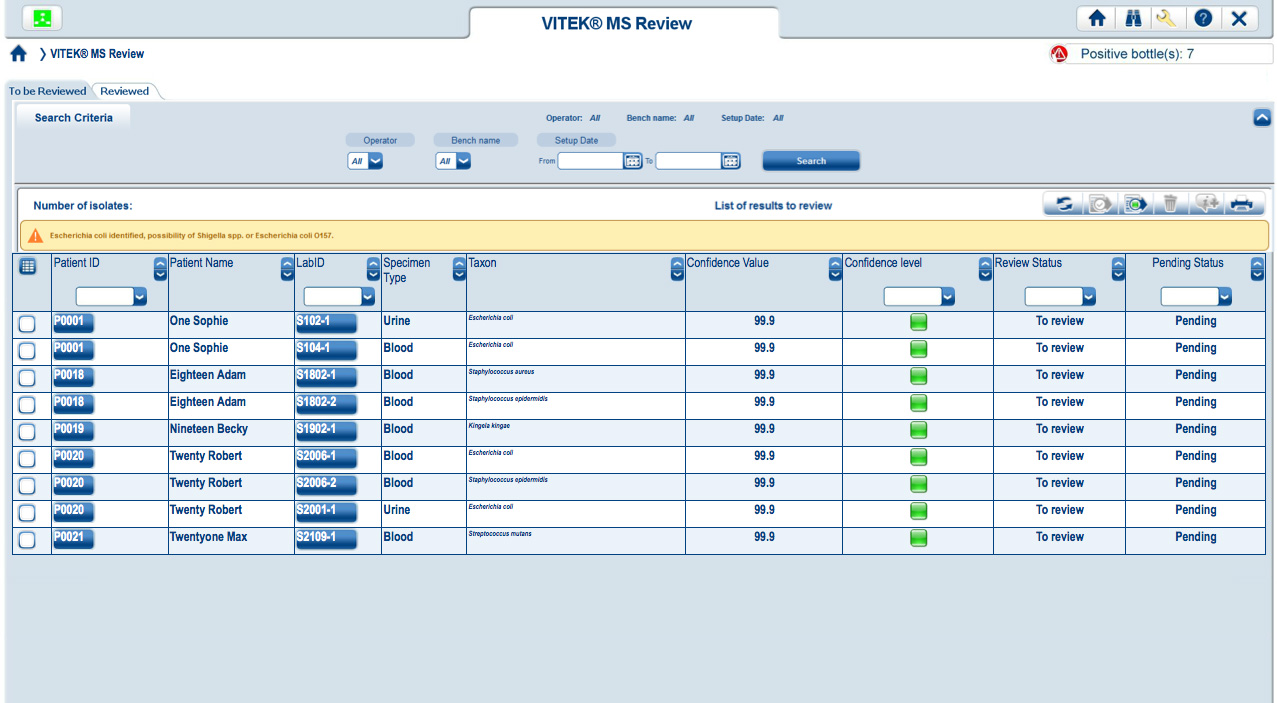
| Easy-to-read results viewed in MYLA |
VITEK SOLUTIONS: Complete Traceability and Flexibility
VITEK MS provides complete traceability and flexibility that contributes to overall lab efficiency and confidence.
- Barcodes on reagents and disposables provide traceability by automatically linking ID with AST results for each isolate.
- Disposable slides:
- Reduce risk of erroneous results from contamination
- Reduce chemical safety hazard issues.
- 3 acquisition areas or groups, each with a calibration spot
- Automatic verification of operation, tuning, and alignment is transparent to the user.
Knowledge Base
bioMérieux's proprietary algorithm – Advanced Spectra Classifier (ASC) - uses a weighted bin matrix to build the Knowledge Base reference strain database. The knowledge base includes data representing 1,316 taxa.
- 207 molds and yeast
- 16 Nocardia
- 39 Mycobacteria
There are on average 40 reference spectra per species covering strain variation, media types, and growth conditions. To create the bin matrix, reference spectra for all species are divided in 1,300 pre-defined bins and bin weights are calculated based on the presence or absence of a peak in a bin and whether this is specific for a certain species compared to all other species in the database.
Once a spectrum is generated from the test sample, the raw data are processed to eliminate baseline values and noise so that significant peaks can be detected. The spectra is then divided in the 1,300 bins and the peak patterns are compared with those for all species in the knowledge base. By computing the sum of the bin weights against all species in the database a relative score is generated and converted to a probability percentage and confidence value. This binning algorithm enables unambiguous identification of an organism with a confidence level greater than 99%.
The approach of analyzing 1,300 data points of each spectrum rather than comparing spectral patterns is very robust for high confidence identification and differentiation of species that have a very closely related protein content and spectral profile. The process enables fast analysis time and improved accuracy over existing methods.
Database Robustness
The VITEK MS is a clinically focused database that contains multiple reference strains from:
- Different regions of the world
- Different specimens
- Different media types and growth conditions to create multiple spectra per species
Medical Value
Delivering outstanding, cost-effective care to patients is highly dependent on a clinician’s ability to access the right information at the right time. VITEK MS supports antimicrobial stewardship programs with microbial identification in minutes and by providing benefits to both laboratory personnel and clinicians.
Results in minutes
- Provides fast and actionable results to initiate optimal therapy
- Broad coverage of clinically significant organisms
Highly accurate results
- Comprehensive database of clinically significant organisms
Rapid results help improve positive patient outcomes
- Reduction in time to optimal antibiotic therapy1
- Helps reduce length of stay, total hospital costs and mortality
Laboratory & Antimicrobial Stewardship Benefits
Benefits for labs
- Ability to identify organisms previously difficult to ID with traditional methodology
- Up to 51.7% decrease in identification cost per isolate2
- Comprehensive & clinically relevant database
- Expanded database of 1,316 organisms including Mycobacteria, Nocardia and Molds
- Now including Brucella species, Candida auris, and Elizabethkingia anophelis
- User-friendly software
- Fully integrated with VITEK® 2 susceptibility testing
Benefits for ASP & clinicians
- Helps reduce time to appropriate therapy3,4
- Helps reduce hospital length of stay5
- High confidence in organism result – highly accurate identifications
References:
2. Tran A, Alby K, Kerr A, Jones M, Gilligan PH. Cost Savings Realized by Implementation of Routine Microbiological Identification by Matrix-Assisted Laser Desorption Ionization–Time of Flight Mass Spectrometry. J Clin Microbiol. 2015;53(8):2473-2479. doi:10.1128/JCM.00833-15
3.
4. Vlek ALM, Bonten MJM, Boel CHE. Direct matrix-assisted laser desorption ionization time-of-flight mass spectrometry improves appropriateness of antibiotic treatment of bacteremia. PLoS One. 2012;7(3): e32589. doi:10.1371/journal.pone.0032589
5. Perez KK, Olsen RJ, Musick WL, et al. Integrating rapid pathogen identification and antimicrobial stewardship significantly decreases hospital costs. Arch Pathol Lab Med. 2013;137(9):1247-1254. doi:10.5858/arpa.2012-0651-OA
Standardized Workflow
The VITEK MS workflow is standardized and efficient, requiring minimal hands-on time.
Bacterial Identification Workflow:
Yeast Identification Workflow:
Efficient workflow
- Spot bacteria and yeasts without extended extraction improves efficiency
- Disposable slides used up to three times
- Efficient processing up to 192 specimens
Workflow Flexibility
The Mycobacterium/Nocardia and mold extraction and inactivation procedures are safe, simple and reliable.1-3
- Multiple media types
- Mycobacteria: BACT/ALERT® MP, Coletsos, MGIT™, Lowenstein-Jensen, Middlebrook 7H10 & 7H11 agar
- Mold: Potato dextrose, Sabouraud dextrose, Sabouraud dextrose with Gentamicin and Chloramphenicol2
- Ability to spot multiple organism types on a single slide (bacteria, fungi, mycobacteria, and Nocardia)
- Extract stability by storage condition
- Mycobacteria extract stability: Frozen (-20°C) for up to 14 days
- Mold extract sample stability: Frozen (-20°C) for up to 14 days
- Slide stability at room temperature – 3 days
Reporting
Easy-to-read, color-coded graphics present the degree of quality and confidence for each ID result.
Results have a strong match and are ready to report Results have low discrimination and require further review Low-quality results with no identification madeVITEK MS connects directly to both VITEK 2 and MYLA® without having to go through the LIS. VITEK MS and VITEK 2 work together, using the same nomenclature, so that VITEK 2 is instantly able to recognize an organism identified by VITEK MS. Through MYLA, VITEK 2 and VITEK MS results are consolidated and reported.
“Mass spec represents a true revolution in microbiology lab automation because it provides a true automated link between the pre-analytic and post-analytic stages. We finally have an option to complete the automation circuit in the microbiology lab for both the testing of samples, but also the reporting of those results.”
Yun F. (Wayne) Wang, MBBS, Ph.D.
Director of Microbiology, Immunology, Molecular Diagnostics
Grady Health System, Atlanta, GA
References:
2. Dunne WM, Doing K, Miller E, et al. Rapid inactivation of Mycobacterium and Nocardia species before identification using matrix-assisted laser desorption ionization–time of flight mass spectrometry. J Clin Microbiol. 2014:52(10):3654-3659. doi:10.1128/JCM.01728-14
3. Totty H, Miller E, Moreno E, Dunne Jr. WM, Deol P. Comparison of mechanical disruption techniques for rapid inactivation of Mycobacterium and Nocardia species before Identification using matrix-assisted laser desorption ionization–time of flight (MALDI-TOF) mass spectrometry. J Clin Microbiol. 2016:54(10):2626-2627. doi:10.1128/JCM.01096-16
4. Refer to FDA 510(k) Clearance Decision Summary.
VITEK MS Technical Data
Laser
- 337 nm nitrogen laser, fixed focus
- Maximum pulse rate - 50 Hz (50 laser shots per second)
- Near normal (on-axis) incidence of the laser beam to the sample
- Laser power and laser aim under software control
Analyzer
- Linear flight tube of 3.9 ft drift length
- Vacuum maintained by two turbomolecular pumps (nominal 250 l/s) with rotary backing
- Beam blanking to deflect unwanted high intensity signals e.g. matrix ions
Dimensions
- Size (w x h x d) – 2 ft 6 in x 6 ft 4 in x 2 ft 10 in, minimum distance to wall at back is 4 in.
- Weight – 805 lbs excluding data system
Installation Requirements
- Power: 4.3 amps @ 220 vac
- Workspace (w x h x d): 8 ft. x 8 ft. x 3 ft. 2 in.
- Floor loading: 123 lbs/ft.2
- Temperature - ambient 18° to 26° C
- Relative humidity - less than 70% non condensing
Components & Disposables
The VITEK MS instrument is a matrix-assisted laser desorption ionization-time of flight mass spectrometer (MALDI-TOF MS) for rapid microorganism identification from clinical cultures.
|
|
The VITEK MS Prep StationThe VITEK MS Prep Station identifies target slides and assigns spots for sample deposits prior to testing on the VITEK MS instrument. In addition, the Prep Station can be used to set up and identify AST cards for testing on the VITEK 2 instrument*. *VITEK 2 instrument and accompanying reagents and disposables are needed for AST testing |
The VITEK MS Acquisition StationThe VITEK MS Acquisition Station is directly connected to the VITEK MS instrument. It displays the status of the tests and the instrument. It acquires the sample spectra data and sends data for analysis to the MYLA server. |
|
|
|
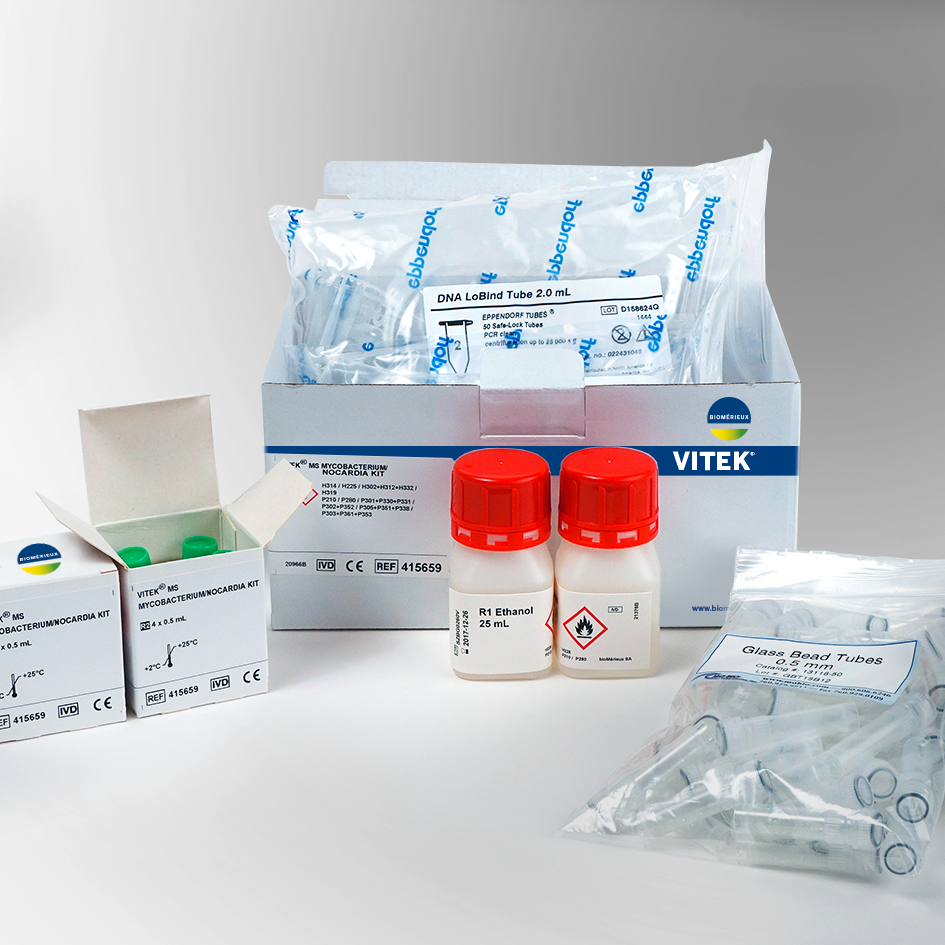
VITEK MS Consumables and ReagentsThe VITEK MS uses disposable, uniquely barcoded target slides for sample deposition. The VITEK MS-CHCA matrix and the VITEK MS-FA are ready to use reagents for the on slide sample preparation. MS-FA is only used to pretreat yeast. The VITEK MS Mycobacterium/Nocardia and Mold reagent kits contain all reagents necessary for simple inactivation and extraction for identification of these organisms on the VITEK MS. |
|
|
MYLA®MYLA is a middleware solution. For VITEK MS, MYLA is connected to both the VITEK MS Prep Station and VITEK MS Acquisition Station providing complete integration and traceability of all individual patient samples and their results. Connectivity to the LIS and other instruments in the laboratory allows all laboratory data to be grouped together and viewed in one place. |
Posters
Flexible Clinical Workflow for Mycobacteria and Nocardia for VITEK® MS (MALDI-TOF): Reliable, Safe, and Efficient for Identification of Positive Patient Samples from Different Media Types
Reproducibility of the New bioMérieux VITEK® MS V3.0 Database for Identification of Moulds, Mycobacterium and Nocardia
Identification of a characterized challenge set of moulds, Mycobacterium and Nocardia strains using the new bioMerieux VITEK® MS V3.0 database
Publications
VITEK MS IVD V3.0
Routine identification of Nocardia species by MALDI-TOF mass spectrometry. Girard V, Mailler S, Polsinelli S, et al.Diag Microbiol Infect Dis. 2017;87(1):7-10. doi:10.1016/j.diagmicrobio.2016.09.024
Identification of mycobacterium spp. and Nocardia spp. from solid and liquid cultures by matrix-assisted laser desorption ionization–time of flight mass spectrometry (MALDI-TOF MS). Girard V, Mailler S, Welker M, et al. Diag Microbiol Infect Dis. 2016;86(3):277-283. doi:10.1016/j.diagmicrobio.2016.07.027
Comparison of sample preparation methods, instrumentation platforms, and contemporary commercial databases for identification of clinically relevant mycobacteria by matrix-assisted laser desorption ionization–time of flight mass spectrometry. Wilen CB, McMullen AR, Burnham C-AD. J Clin Microbiol. 53(7):2308–2315. doi:10.1128/JCM.00567-15
Matrix-assisted laser desorption ionization-time of flight (MALDI-TOF) mass spectrometry using the VITEK® MS system for rapid and accurate identification of dermatophytes on solid cultures. De Respinis S, Monnin V, Girard V, et al. J Clin Microbiol. 2014:52(12):4286-4292. doi:10.1128/JCM.02199-14
Rapid inactivation of mycobacterium and Nocardia species before identification using MALDI-TOF mass spectrometry. Dunne WM Jr, Doing K, Miller E, et al. J Clin Microbiol. 2014:52(10):3654-3659. doi:10.1128/JCM.01728-14
VITEK MS IVD V2.0 (In Vitro Diagnostic Applications)
Matrix-assisted laser desorption ionization time of flight mass spectrometry (MALDI-TOF MS) for rapid identification of micro-organisms in the routine clinical microbiology laboratory. Wattal C, Oberoi JK, Goel N, et al. Eur J Clin Microbiol Infect Dis. 2017:36(5):807-812. doi:10.1007/s10096-016-2864-9
Comparison of VITEK® MS and MALDI Biotyper® for identification of Actinomycetaceae of clinical importance.Ferrand J, Hochard H, Girard V, et al. J Clin Microbiol. 2016:54(3):782-784. doi:10.1128/JCM.02758-15
MALDI-TOF mass spectrometry for differentiation between Streptococcus pneumoniae and Streptococcus pseudopneumoniae. van Prehn J, van Veen S, Schelfaut J, Wessels E. Diagn Microbiol Infect Dis. 2016;85(1):9-11. doi:10.1016/j.diagmicrobio.2016.01.012
Using matrix-assisted laser desorption ionization-time of flight (MALDI-TOF) complemented with selected 16S rRNA and gyrB genes sequencing to practically identify clinical important viridans group streptococci (VGS). Zhou M, Yang Q, Kudinha T, et al. Front Microbiol. 2016;7:1328. doi:10.3389/fmicb.2016.01328
Assessment of reproducibility of matrix-assisted laser desorption ionization–time of flight mass spectrometry for bacterial and yeast identification. Westblade LF, Garner OB, MacDonald K, et al. J Clin Microbiol. 2015;53(7):2349–2352. doi:10.1128/JCM.00187-15
Comparative evaluation of two matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS) systems, VITEK® MS and Microflex LT, for the identification of Gram- positive cocci routinely isolated in clinical microbiology laboratories. Lee M, Chung HS, Moon HW, Lee SH, Lee K. J Microbiol Methods. 2015;113:13-15. doi:10.1016/j.mimet.2015.03.020
Performances and reliability of Bruker Microflex LT and VITEK MS MALDI-TOF mass spectrometry systems for the identification of clinical microorganisms. Bilecen K, Yaman G, Ciftci U, Laleli YR. Biomed Res Int. 2015;Article ID 516410:18 pages. doi:10.1155/2015/516410
Comparison and optimization of two MALDI-TOF MS platforms for the identification of medically relevant yeast species.Pence MA, McElvania TeKippe E, Wallace MA, Burnham CA. Eur J Clin Microbiol Infect Dis. 2014;33(10):1703-12. doi:10.1007/s10096-014-2115-xhttps://doi.org/10.1007/s10096-014-2115-x
MULTICENTER EVALUATIONS OF THE VITEK MS IVD V2.0 SYSTEM
Multicenter validation of the VITEK® MS v2.0 MALDI-TOF mass spectrometry system for the identification of fastidious gram-negative bacteria. Branda JA, Rychert J, Burnham CA, et al. Diag Microbiol Infect Dis. 2014;78(2):129-31. doi:10.1016/j.diagmicrobio.2013.08.013
Multicenter evaluation of the VITEK® MS system for mass spectrometric identification of non-Enterobacteriaceae Gram-negative bacilli.Manji R, Bythrow M, Branda J, et al. Eur J Clin Microbiol Infect Dis. 2014;33(3):337-346. doi:10.1007/s10096-013-1961-2
Multi-centre evaluation of mass spectrometric identification of anaerobic bacteria using the VITEK® MS system. Garner O, Mochon A, Branda J, et al. Clin Microbiol Infect. 2014;20(4):335-339. doi:10.1111/1469-0691.12317
Multicenter evaluation of the VITEK MS matrix-assisted laser desorption ionization-time of flight mass spectrometry system for identification of Gram-positive aerobic bacteria. Rychert J, Burnham CA, Bythrow M, et al. J Clin Microbiol. 2013:51(7):2225-2231. doi:10.1128/JCM.00682-13
Multicenter study evaluating the VITEK MS system for identification of medically important yeasts. Westblade LF, Jennemann R, Branda JA, et al. J Clin Microbiol. 2013;51(7):2267-2272. doi:10.1128/JCM.00680-13
Identification of Enterobacteriaceae by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry using the VITEK® MS system. Richter SS, Sercia L, Branda JA, et al. Eur J Clin Microbiol Infect Dis. 2013;32(12):1571-1578. doi:10.1007/s10096-013-1912-y